A New Kind of Non-Acoustic Speech Acquisition Method Based on Millimeter Waveradar
Sheng Li,
Ying Tian,
Guohua Lu,
Yang Zhang,
Hui Jun Xue,
Jian-Qi Wang and
Xi-Jing Jing
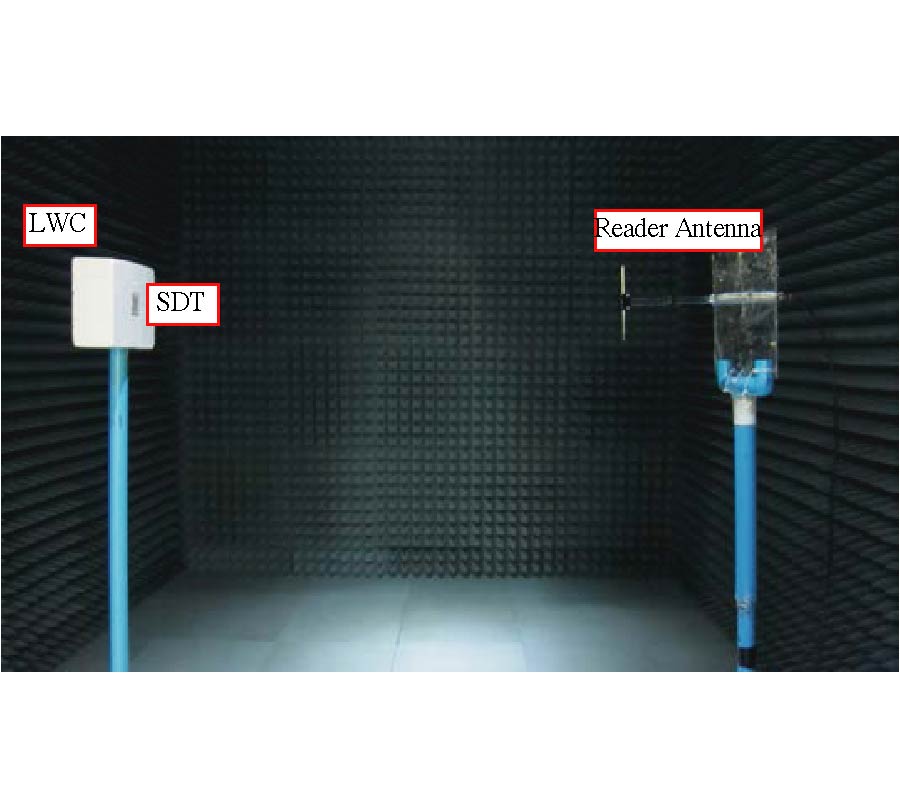
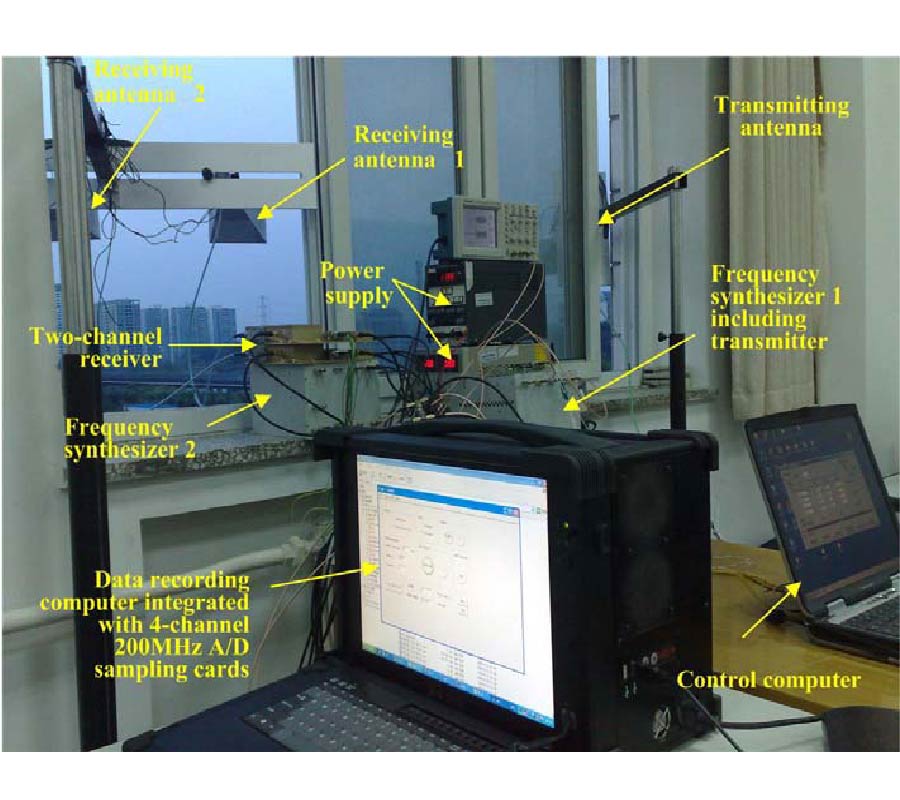
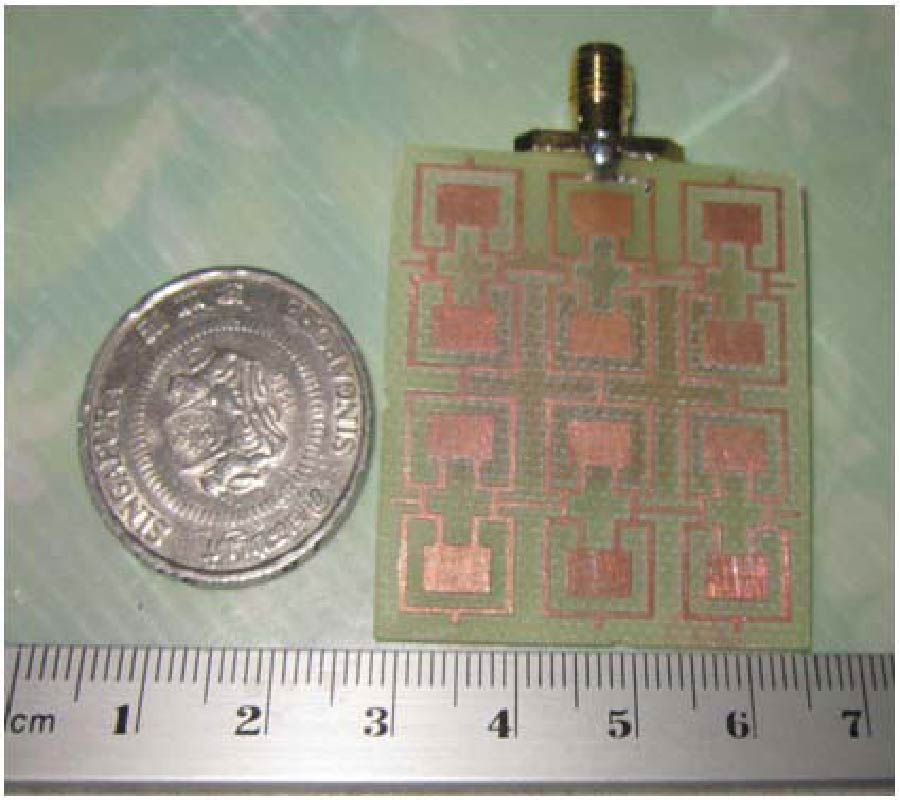
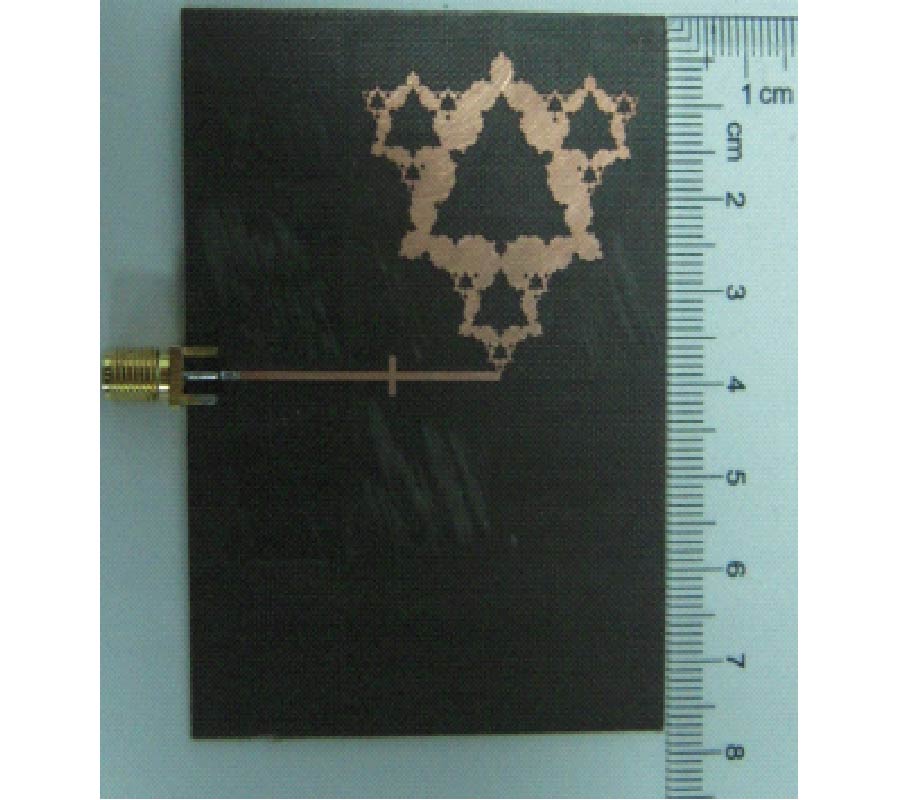
Air is not the only medium that can spread and can be used to detect speech. In our previous paper, another valuable medium - millimeter wave (MMW) was introduced to develop a new kind of speech acquisition technique [Li et al., Progress In Electromagnetics Research B, 9, 199-214, 2008]. Because of the special features of the MMW radar, this speech acquisition method may provide some exciting possibilities for a wide range of applications. In the proposed study, we have designed a new kind of speech acquisition radar system. The super-heterodyne receiver was used in the new system so that to mitigate the severe DC offset problem and the associated 1/f noise at baseband. Furthermore, in order to decrease the harmonic noise, electro-circuit noise, and ambient noise which were combined in the MMW detected speech, an adaptive wavelet packet entropy algorithm is also proposed in this study, which incorporates the wavelet packet entropy based voice/unvoiced radar speech adaptive detection method and the human ear perception properties in a wavelet packet time-scale adaptation speech enhancement process. The performance of the proposed method is evaluated objectively by signal-to-noise ratio and subjectively by mean-opinion-score. The results confirm that the proposed method offers improved effects over other traditional speech enhancement methods for MMW radar speech.